Codon recoding
What it is:
Making the genetic code a little less universal
Imagine that you are an explorer traveling from your homeland to uncharted territories long ago. Now imagine that every new person you meet in these distant lands, from sea-faring cultures on one side of the world to jungle tribes on the other, greeted you in the language you grew up speaking, perfectly fluently with no accent at all. This would seem remarkably unlikely; in fact, if you travel the world even today you will encounter people speaking thousands of languages, all different from each other.
Yet in genetics there is a single universal language. Almost every organism we have ever encountered uses the exact same genetic code. That is, DNA from almost any organism on Earth can be placed in almost any other organism, and it will be read the exact same way. It’s as if you could pick up any book ever written from any library on Earth and read it in perfect modern English. For this reason, we call the genetic code universal. In every organism, the four nucleotides of DNA, adenine (A), thymine (T), guanine (G) and cytosine (C), are read in groups of three called codons, and in almost every organism each specific group of three nucleotides spells out the same protein-coding instruction. ATG codes for the amino acid methionine, TCG codes for serine, GGA for glycine, etc.
That every organism uses the same code has been a huge benefit to scientists. It means that we can, just by looking at the DNA of a previously unknown organism, predict exactly what proteins it will make. It also means that we can take DNA from one organism and place it in a completely unrelated organism and be able to predict how it will function. But scientists are now asking, what if we could create an organism that used a slightly different code? Does the fact that there is only one code limit our applications of biotechnology?
How it works:
Taking advantage of redundancy
There are 64 different ways to arrange the four nucleotides (A, T, C, and G) into groups of three. But there are only 20 amino acids. This allows for redundancy in the genetic code – there is more than one way to code for the same amino acid. There are six different three-nucleotide combinations that code for the amino acid leucine, four ways to code for alanine, and so on. This redundancy means that, theoretically, you could change a codon to code for something new or even eliminate it altogether. Life could go on with only five ways to code for leucine. The problem is that in cell that is using all of its possible codons, simply getting rid of one would make a lot of genes unreadable and kill the cell.
Conceptually, recoding the genome is fairly simple. First, pick a codon that is redundant.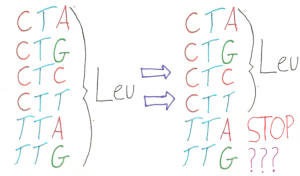 Second, change every place in the genome where that codon is found to a different codon that codes for the same amino acid. Third, engineer the cell to use that original codon in a new way or not at all. In practice, it is much more complex. The second step of this process requires potentially thousands of very specific changes throughout a genome. Even using new genetic engineering technology such as CRISPR/Cas9, this is too large and specific a genetic engineering task. The creation of synthetic genomes has given scientists a new approach to this problem. Now, instead of trying to make thousands of changes to a functional genome while inside a living organism, scientist can synthesize the new DNA sequence on a machine with all the changes they want already in it. This new DNA is then added into existing bacteria through recombination. This process of engineering organisms by using recombination to introduce large sequences of synthesized DNA has been given the name “recombineering”.
Second, change every place in the genome where that codon is found to a different codon that codes for the same amino acid. Third, engineer the cell to use that original codon in a new way or not at all. In practice, it is much more complex. The second step of this process requires potentially thousands of very specific changes throughout a genome. Even using new genetic engineering technology such as CRISPR/Cas9, this is too large and specific a genetic engineering task. The creation of synthetic genomes has given scientists a new approach to this problem. Now, instead of trying to make thousands of changes to a functional genome while inside a living organism, scientist can synthesize the new DNA sequence on a machine with all the changes they want already in it. This new DNA is then added into existing bacteria through recombination. This process of engineering organisms by using recombination to introduce large sequences of synthesized DNA has been given the name “recombineering”.
This process has been used to demonstrate the principle of codon recoding in E. coli by a team at Harvard University led by George Church. In 2013, the team announced that they had recoded every “TAG” stop codon in the E. coli genome to a synonymous “TAA” stop codon. In 2016, the team announced that they had recoded the entire E. coli genome in 55 different segments, in each, entirely eliminating the use of seven different codons. To do so required 62,214 individual genetic changes. They have yet to combine all 55 segments into a single functional genome.
The future:
Virus immunity and new biochemistry
When a virus releases its genetic material into a cell, it relies on the host cell’s ribosomes and tRNAs to make the proteins the virus needs to reproduce. Because there is no difference between virus genetic code and the host cell genetic code, the viral instructions are read just as if they were from the cell’s own DNA. In a recoded organism, when a virus invaded the cell, its code would be different from that of the host. As the virus tried to hijack the cell’s machinery, the instructions spelled out by its genetic material would no longer make any sense. It would be as if someone tried to copy your homework, but you had written every few words in your own secret made up language. It doesn’t matter who tried to copy it, it wouldn’t make any sense. In essence, the cell will have been made immune to all viruses, even ones we have not discovered yet.

When a codon is no longer used it could also be repurposed. Scientists could program unused codons to code for new, not naturally occurring amino acids, creating proteins that could not previously exist in biology. Some scientist are already starting to test ways to incorporate these new amino acids into functional proteins. These techniques are still in development, so nobody knows for sure what doors this new biochemistry will open for scientists in the future. But if the past is any indication, when new genetic technologies are created, scientists will find uses for them.
Questions
Review:
- What do scientists mean when they say the genetic code is universal?
- Explain why there is redundancy in the genetic code.
- What would be wrong with just eliminating the use of a codon without recoding it first?
- Why do scientists need to use synthetic DNA and recombineering instead of other genetic engineering techniques when recoding codons?
- Why are viruses especially susceptible to codon recoding as a cellular defense mechanism?
Critical thinking:
- The universal genetic code is actually not quite universal. There are several examples of genomes that have small changes to the universal genetic code. These include different mitochondrial genomes and some other, typically small, genomes. It is believed that these alternative codes evolved naturally from the universal code following the same three basic steps outlined above. These changes would have happened randomly, by chance. Why do you think these naturally recoded genomes are typically found in very small genomes and not larger ones?
- Do you think codon recoding could be used for humans, making us all resistant to every virus?
Discussion:
- Some people suggest using codon recoding to protect against dangerous genetically modified organisms escaping from a laboratory?
Answer key: Available to teachers upon request: dnadots@minipcr.com
